Safe storage of large data sets within the DNA in living cells has become practical, says S.Ananthanarayanan.
In the days when information was written down and letters of the alphabet had to be recognised by the eye, records could not be packed closer than what could be easily seen. But in the era of computers, devices can detect ‘0s’ and ‘1s’ that are written very close together. Data that is converted to the ‘0 and 1’, or the binary form, can hence be stored in great quantity, in media like magnetic tape, hard disks or pen drives.
Even these, however, are proving insufficient, as the data needs of applications have become very large indeed. Another difficulty with the existing forms of storage is that records degrade with age or need to be transferred to new media when there are changes in technology.
Being able to store data in the DNA of cells has hence been viewed as a solution made to order. This is because the DNA molecule has both great capacity and is exceedingly hardy. We know DNA last long because DNA from scraps of once living tissue, which are discovered in archaeological remains, are intact for analysis!
Seth L. Shipman, Jeff Nivala, Jeffrey D. Macklis and George M. Church from Harvard Medical School and Harvard University report in the journal, Nature, that they have scaled up a method of using native, biological tools to insert encoded data, which has been recorded in a synthetic molecule, into the DNA of living cells. In a proof of principle demonstration, a movie clip of a galloping horse has been secured in a single living cell.
Shipman and colleagues explain in the paper that the trick involves two steps. The first is to encode the information in the form of a sequence of units like the ones found in the DNA molecule. And the second step is to snip the DNA molecule at a specified place, to insert the sequence that has the information.
These DNA-like portions that are inserted into DNA are molecules that are built in the same way DNA. Hence, like DNA, they consist, of a chain, or backbone of units, called nucleotides or bases, as shown in the picture, with ‘side chains’ of four kinds of molecular groups, along the length of the DNA backbone. The DNA, and the shorter variety, which are called oligonucleotides, carry information, as coded by the sequence of the four kinds of side chain groups. These groups have the names, C, G, A and T and in the DNA, groups of three consecutive bases, each with one of four forms of side chain, code for twenty amino acids, which are the building blocks of proteins. Series of triads thus code for series of amino acids and hence for different proteins.
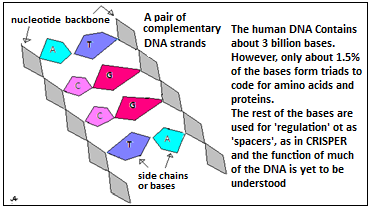
Just as the DNA codes for proteins, the shorter oligonucleotides can code digital information. In computers, groups of eight ‘bits’, or binary units, code for 28=256 characters. In DNA, a group of three bases, which can take four forms each, can have 43=64 forms (but only twenty amino acids are coded, as some acids have alternate codes, for safety). Oligonucleotides, which have the same structure, can hence be used to form codes to represent strings of text characters, numbers or pixel values. In practice, it is found that more than three consecutive bases with the same side chain creates instability in oligonucleotides. The side chain, ‘G’, is hence reserved to break series of more than three identical bases and only the remaining three side chains are there for coding.
The Harvard paper describes a scheme of coding in the synthesised segments, where the pixels in an image of a hand, as shown in the picture, are represented by the side chains along the length of bases. As there is a limit to how long the segments can be, coding is done over more than one set of synthetic bases. To identify which pixels a segment represents, a part of the segment is used for identification. After this and other overheads, only 28 bases were left in the segments for coding pixels.
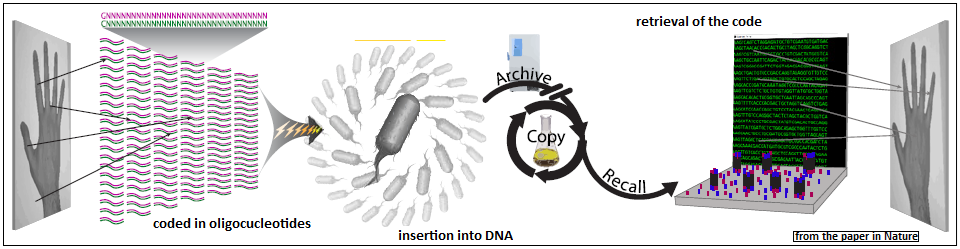
Entering the DNA
The second step in the process was to insert the set of oligonucleotides into the DNA of a living cell. This was done with the help of a feature of real cells and which has been perfected for artificial manipulation of the DNA. The mechanism by which cells gain immunity against virus attacks is that when a virus attacks a cell, the cell copies signature portions of the viral DNA into a portion of its own DNA. This part of the cell DNA, called CRISPR, for Clusters of Regularly Interspaced Short Palindromic Repeats, is not used for the cell’s normal functions, but if the virus should attack again, the cell’s defences have a copy of unique parts of the virus DNA. Another part of the cell DNA, called CAS genes, for CRISPR Associated genes, now use this information to go out and snip, or divide, the viral DNA at the place identified.
This method, native to cells, was then turned around to cut the DNA of a cell itself, at specific places. The cut ends would then spontaneously join, and in the process, there could be repair of defective DNA or the insertion of a portion to add to the genes that are present in normal DNA. The method, called CRISPER-CAS9, has taken genetic engineering by storm and has set in motion great new work and advances in the field.
The Harvard group made use of CRISPR-CAS9 to insert the panel of oligonucleotides, which had been fashioned to record digital data, into the DNA of living cells. The DNA would then be efficiently replicated, in the process of cell division, and preserved, with economy and security! There are now efficient methods of reading the sequence of side chains along the length of the bases of DNA. These techniques enable the information coded in the inserted segments to be read, or retrieved, for use.

That the technique works was demonstrated by recording a panel of a pictures of a galloping horse, by Eadweard Muybridge, a celebrated 19th century photographer of people and animals in motion. The pictures, as shown, could be recorded and retrieved with good preservation of quality, and displaying the pictures in succession created a motion picture of the horse in action, all recorded inside the DNA of a living cell!
------------------------------------------------------------------------------------------